Spectracular
Welcome to Spectracular, the online tool for minimizing spectra overlap in multicolor flow cytometry panels.

|
What is Spectracular?
Spectracular is a computational tool to support selection of fluorochromes for multicolor flow cytometry panels. Specifically, Spectracular finds the optimal (or near-optimal) combination of fluorochromes from a user-defined pool of fluorochromes to choose from.
How to use Spectracular
In order to use spectracular, you need to know how many fluorochromes you would like in your panel, out of which pool of fluorochromes you want to choose, and the laser configuration of your flow cytometer.
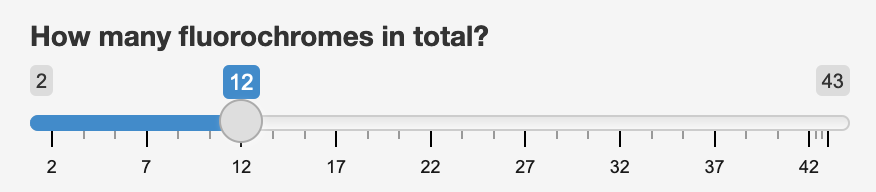
|
Note that as a default, you can choose between 2 and 43 - this is because the default pool of fluorochromes that Spectracular consideres consist of 44 commonly used fluorochromes. The slider will automatically increase and decrease in range as you add or remove fluorochromes to consider.

|
This reveals the full list of 133 fluorochromes to choose from. While it will not change the runtime of the algorithm much to include all of them, we recommend selecting a reasonable subset based on what fluorochromes are available for the antibodies you intend to use.
Immediately below that, is a dark blue “Fluorochromes to include”" button. Clicking this will reveal a list consisting of the fluorochromes selected above. From these, you may select a subset of no more than k fluorochromes that will be included in your solution, regardless of their overlap. This option allows users to force Spectracular to consider fluorochromes that they may want to use for reasons beyond overlap.
|
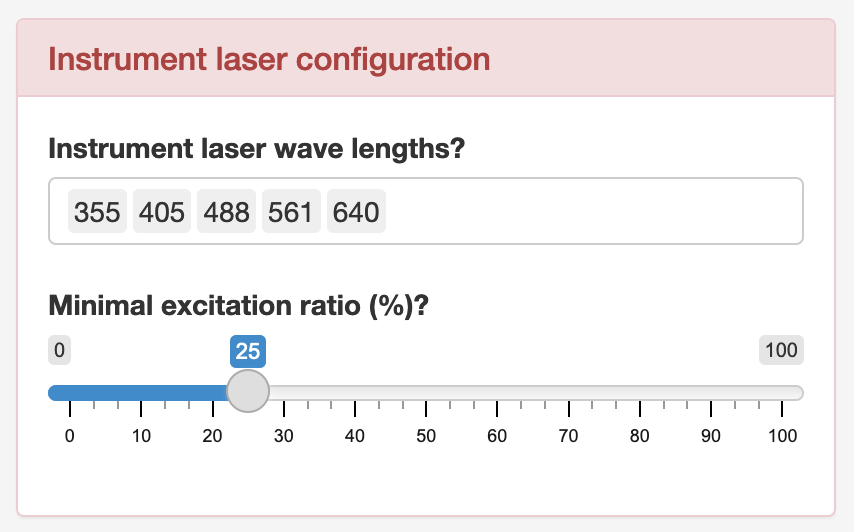
Lastly, clicking the red “Instrument laser configuration” button reveals the following dialog box:
|
Lasers are added by typing the wavelength (in positive integers between 300 and 900) and hitting enter. Lasers can be removed by placing the cursor in front of a laser and hitting delete. The minimal excitation ratio, is a threshold for including fluorochromes in the final solution. A minimal excitation ratio of 25% means that in order for a fluorochrome to be considered for the final solution, it must be excited to at least 25% of the theoretical maximum excitation. A fluorochrome will be assumed to be excited by the laser which leads to the highest emission signal.
How does Spectracular work?
Calculating the total overlap of every combination of k fluorochromes out of a pool of n, is a combinatorial problem with the complexity:This means that if you wish to find the optimal combination of, say, 20 fluorochromes out of 133 fluorochromes currently available in spectracular, we would need to calculate the sum overlap coefficient of:
This is a lot of combinations to calculate. For comparison, there are an estimated \(7.5\times10^{18}\) grains of sand on earth and somewhere between \(10^{22}\) and \(10^{24}\) stars in the observable universe. Given that the computational problem is NP hard (which roughly means that the effort required to solve the problem grows exponentially with the size of \(N\)), this number of combinations is essentially not possibly to calculate using a brute force approach. Spectracular approximates the optimal solution by intelligently reducing the combinatorial complexity (the precise algorithm is detailed in our bioRxiv paper).
Problems?
Questions, comments, or issues? Please contact us (see contact info at the bottom of this page).
Contact